- From: William Bug <William.Bug@DrexelMed.edu>
- Date: Sun, 17 Dec 2006 23:26:41 -0500
- To: Joanne Luciano <jluciano@predmed.com>
- Cc: kc28 <kei.cheung@yale.edu>, Alan Ruttenberg <alanruttenberg@gmail.com>, W3C HCLSIG <public-semweb-lifesci@w3.org>
- Message-Id: <B47572C7-F364-4CF9-9E3D-A6E737F9FC8C@DrexelMed.edu>
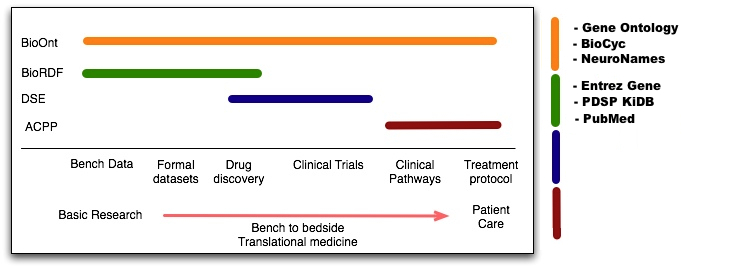
Hi Joanne, I like what you are trying to achieve in this figure, and I think you do it well in a very small amount of space. I've added just a quick "key" to the right-hand side, to give readers a more concrete sense of what community resources the 4 Task Forces have been focused on integrating. This should obviously not be exhaustive, just examples to give folks a concrete sense of what each task force has been examining. I provided examples for BioONT and BioRDF drawn from the paper. I'd leave it to those who know better than I what public resources fall under the purview of DSE & ACPP. Just my $0.02. Cheers, Bill  On Dec 17, 2006, at 10:10 PM, Joanne Luciano wrote: > > Please provide feedback on the figure too. > > Joanne > > On Dec 17, 2006, at 2:53 PM, kc28 wrote: > >> >> Hi Joanne, >> >> I'll work with Alan and others based on your suggestions. >> >> Thanks, >> >> -Kei >> >> Joanne Luciano wrote: >> >>> Hi Kei, >>> >>> I made my edits on the google page this morning, so this becomes >>> a belated request for deadline extension. >>> >>> Included here: >>> >>> 1.3.2. Standards as Applications of Semantic Web Technologies >>> (Joanne) >>> >>> >>> There is a tacit assumption within the Semantic Web community >>> that every type of data sets and ontologies will be >>> interoperable. The reality is that a multitude of different >>> conceptualisations and formal representation of data exist. A >>> key part of the Health and Life Sciences SIG's vision is that >>> standards need to be established and accepted sufficiently >>> widely within the community so as to allow that necessary >>> interoperability between data sets, application and workflows. A >>> relatively early effort in this direction was the BioPAX >>> initiative [REFS] which aimed to provide a common framework for >>> the disparate data sources and multiple conceptualisations of >>> cellular pathways. Its major success was in increasing public >>> awareness and persuading a community of researchers that the >>> integration of data was possible and would result in major >>> research and discovery advantages. It highlighted the multiple >>> conceptualizations existing within the domain of cellular >>> pathways and made people aware of the significant investment >>> needed to achieve interoperability at not only a syntactic level >>> but also at a semantic one. A considered critique of the effort >>> with suggestions of the way forward is presented in [REF to >>> Luciano and Stevens] in this issue. >>> >>> In the medium term future, the efforts for standardization need >>> a much more careful analysis of the user requirements so as to >>> be able to get the end users (wet lab scientists) to buy into >>> the approach and its benefits. A greater understanding is needed >>> both of the conceptualisations needed which will be reflected in >>> a number of ontologies and the boundary which lies between what >>> is possible and useful in OWL based ontologies given the >>> capabilities of the current generation of reasoners, and on the >>> other hand the ambitions and needs of clinical researchers to >>> integrate and interpret their data. >>> >>> The HCLSIG has an interesting role in the SW community because >>> potentially there is a far greater commitment to resourse >>> curation and the need for standard formats and data integration, >>> thus making it a much more likely candidate for being a poster >>> child for the success of Semantic Web technologies. >>> >>> Added to Section 2: >>> >>> >>> The following figure shows the respective relationship between >>> the different task forces and their role in the bench to bedside >>> vision. >>> >>> >>> -------------------------------------------------------------------- >>> ---- >>> >>> >>> and added my contributions to the section at the bottom: >>> >>> >>> JL Joanne Luciano, Havard Medical School, Universtity of >>> Manchester (UK), BioPathways Consortium, BioPAX >>> >>> Affiliation: >>> >>> JL Joanne Luciano, Havard Medical School, Universtity of >>> Manchester (UK), BioPathways Consortium, BioPAX >>> >>> To Acknowledgment >>> >>> (after Kei's) JL was supported by NSF grant IIS-0542041. >>> >>> ---Joanne >>> >>> >>> On Dec 15, 2006, at 10:58 AM, Kei Cheung wrote: >>> >>>> >>>> Just want to add that the google doc editing deadline is 12:00 >>>> am (mid-night EST), Dec 15. If people still need a little bit >>>> more time, please let us know as soon as possible. We might be >>>> able to extend it to 12:00 noon, Saturday Dec 16 (but not >>>> anymore since the deadline is very close). >>>> >>>> Thanks, >>>> >>>> -Kei >>>> >>>> Alan Ruttenberg wrote: >>>> >>>>> >>>>> Correction: Helen, Vipul will send whatever they haven't >>>>> integrated into the googledoc to Alan over the weekend for >>>>> integration. Alan will edit and send to Kei at weekend's end. >>>>> >>>>> -Alan >>>>> >>>>> On Dec 14, 2006, at 6:14 PM, Eric Neumann wrote: >>>>> >>>>>> [NEW] ACTION: Content will be moved off from googleDoc to >>>>>> Word COB Dec 15; to be serially edited by Alan, Helen, >>>>>> Vipul, and returned to Kei Dec 20. [recorded in http:// >>>>>> www.w3.org/ 2006/12/14-hcls- minutes.html#action01 <http:// >>>>>> www.w3.org/ 2006/12/14-hcls- minutes.html#action01> ] >>>>>> >>>>> >>>>> >>>> >>>> >>>> >>>> >>> >>> Joanne Luciano, PhD >>> Predictive Medicine, Inc. >>> 45 Orchard Street >>> Belmont MA 02478-3008 >>> Email: jluciano@predmed.com >>> >>> >>> >> >> >> > > Joanne Luciano, PhD > Predictive Medicine, Inc. > 45 Orchard Street > Belmont MA 02478-3008 > Email: jluciano@predmed.com > > > > Bill Bug Senior Research Analyst/Ontological Engineer Laboratory for Bioimaging & Anatomical Informatics www.neuroterrain.org Department of Neurobiology & Anatomy Drexel University College of Medicine 2900 Queen Lane Philadelphia, PA 19129 215 991 8430 (ph) 610 457 0443 (mobile) 215 843 9367 (fax) Please Note: I now have a new email - William.Bug@DrexelMed.edu
Attachments
- text/html attachment: stored
- image/jpeg attachment: HCLSIG_TF-overlap-BBedit.jpg
- text/html attachment: stored
Received on Monday, 18 December 2006 04:27:05 UTC