- From: James Malone <malone@ebi.ac.uk>
- Date: Wed, 30 Nov 2011 17:23:09 +0000
- To: HCLS <public-semweb-lifesci@w3.org>
- CC: Marco Brandizi <brandizi@ebi.ac.uk>
- Message-ID: <4ED6667D.7@ebi.ac.uk>
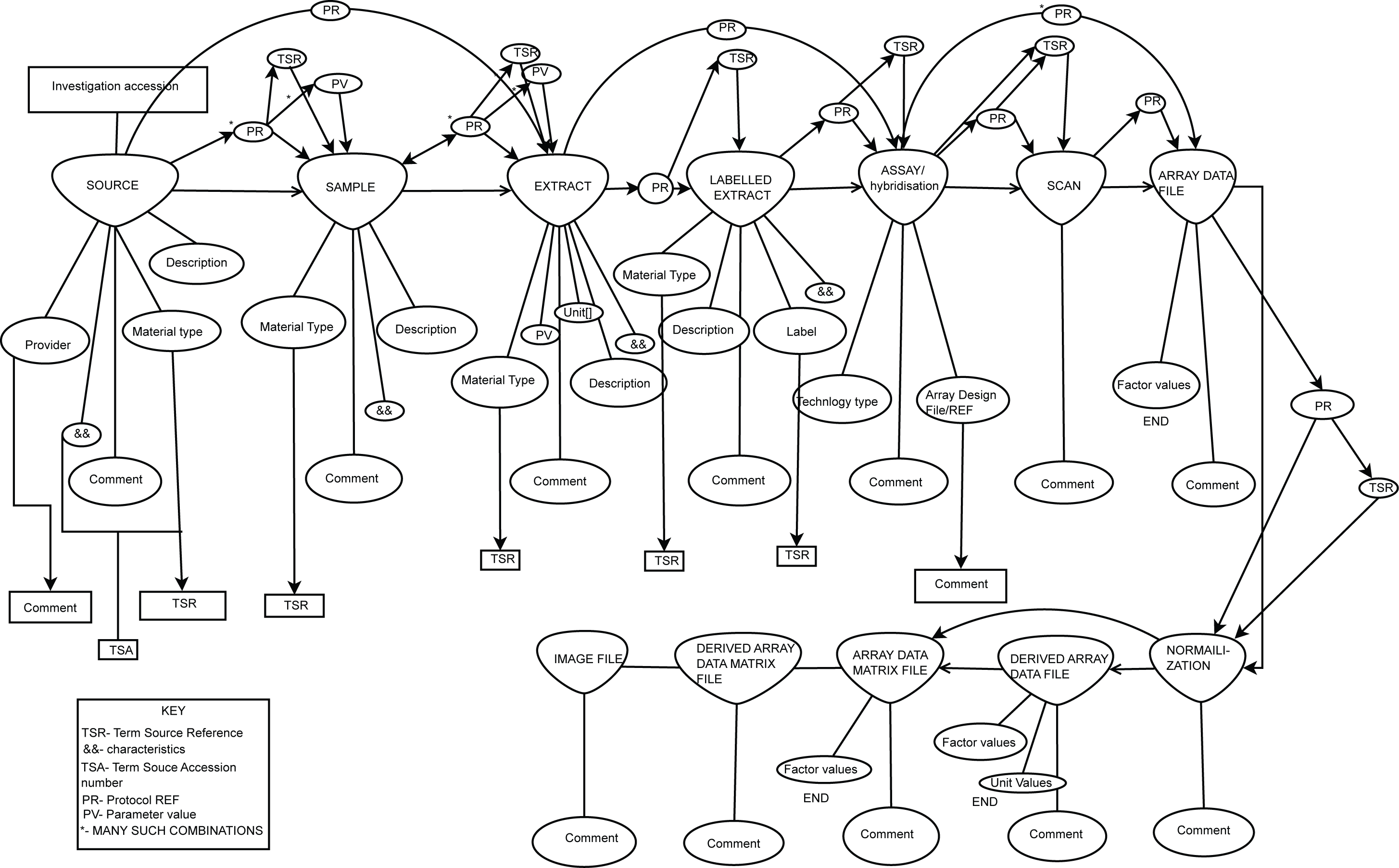
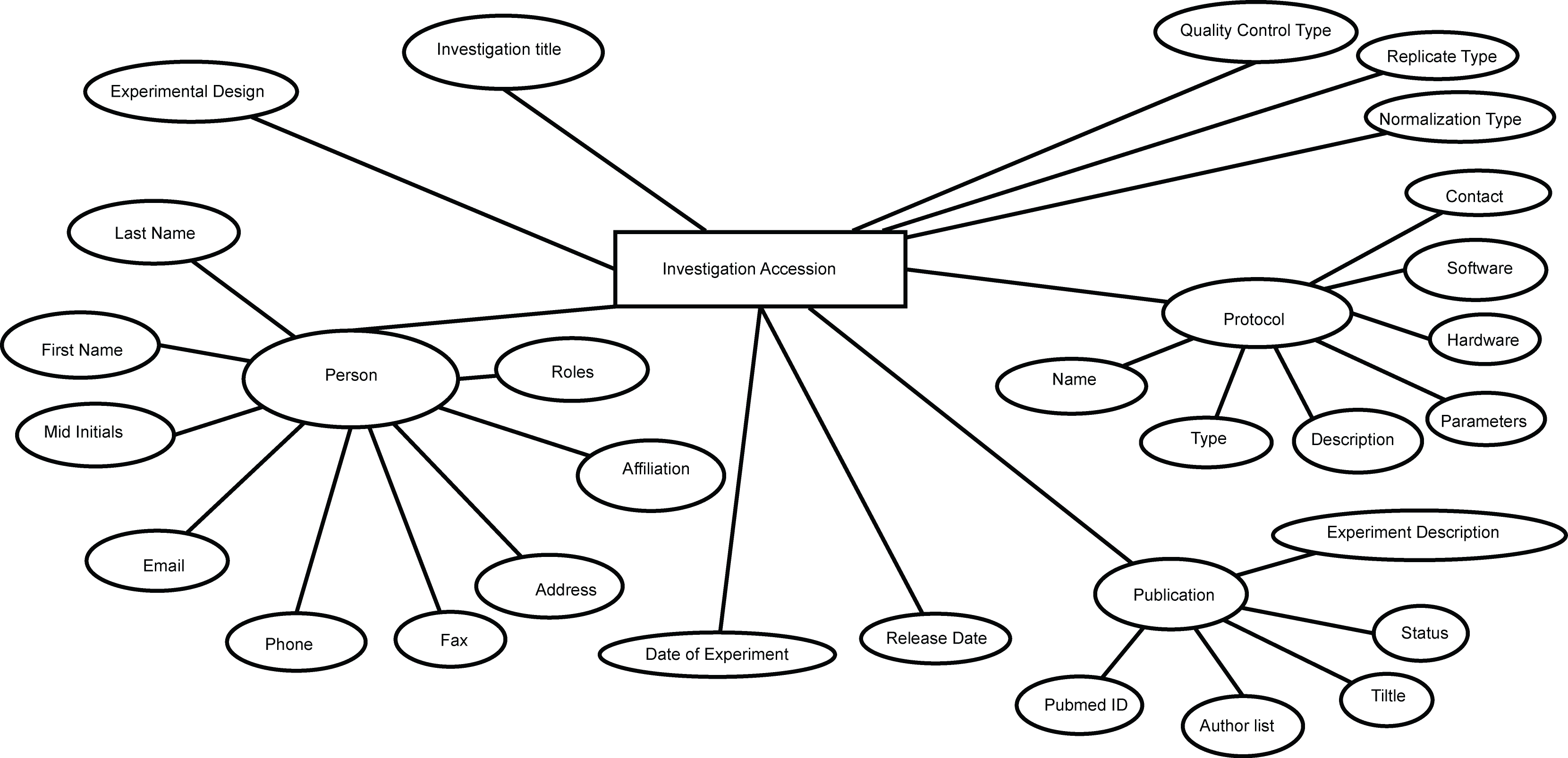
All, I had promised to send something on regarding the work our student (Drashtti Vasant) had been doing on producing RDF from our MAGE-TAB experiments in ArrayExpress. So here are some bits. We started this a couple of months back before a model was sent around by Jim or Michael (apologies, not sure who it was exactly - for whatever reason I can not find the email and diagram that was sent round - could someone resend it please!). Interesting to see the things that align/different, I seem to recall they aren't a million miles off. Anyway, I've attached the graphs we used as our baseline and the table of nodes we included as rdf and the triples we formed in this rdf - note we do not include everything in the graphs attached. For factor values, we ran our Zooma term matching across the values to try and mine these against matches in EFO and for those that we matched we formed triples (e.g. liver, cancer). For those we couldn't match automatically we added some default high level triples (which are less useful) just to say this has some factor values and we captured it as a literal text string. I'm simplifying a bit as it was a bit more complicated but more or less that is what we did. As well as the triples in the doc I've attached we also added some "convenience" triples to each of these nodes directly to the experiment using part_of relation so you can do simple queries on the experiment without having to traverse the whole graph. The store is sitting on an internal server at the moment but we may open it up if there is any interest in using it. Otherwise, we have some internal (interesting) stuff we're doing with it as well expanding and refining what we have. We focused on things we wanted to ask questions about explicitly based on a set of competency questions we had formed. It's not perfect but I'm of the school of release early, often, refine. Cheers, James PS credit for work is to Drashtti Vasant who did most of this implementation, myself and Tony Burdett supervised with some guidance on RDF matters from Marco Brandizi. -- European Bioinformatics Institute, Wellcome Trust Genome Campus, Cambridge, CB10 1SD, United Kingdom Tel: + 44 (0) 1223 494 676 Fax: + 44 (0) 1223 494 468
Attachments
- image/png attachment: sdrf.png
- image/png attachment: idf.png
- application/msword attachment: ontology_classes.doc
- application/msword attachment: triples.doc
Received on Wednesday, 30 November 2011 17:24:13 UTC